Our lab is primarily interested in the phylum Neocallimastigomycota, the fungal phylum of anaerobic gut fungi (AGF). AGF play a role in plant biomass degradation in many herbivorous animals. The lab is active in investigating the ecology, metabolic capabilities, and genomics of the Neocallimastigomycota.
We are also interested in the metagenomics and metatranscriptomics of anoxic environments. In addition to this, we are exploring the causes of algal blooms in freshwater lakes.
In summary, our research is active on several fronts:
- Isolation, characterization, and genomic sequencing of anaerobic gut fungi
- Microbial ecology of anoxic environments: metagenomics and metatranscriptomics.
- Investigating the source of harmful algal blooms in freshwater lakes
Projects
Select a project to view more information.
Anaerobic Fungi
Isolation and characterization of novel anaerobic fungal genera. Anaerobic gut fungi (AGF) are hard to isolate and maintain. They are oxygen sensitive, prone to senescence, and have very narrow physiological optima. In addition to this, procedures for their long-term storage are often unreliable. Nevertheless, our laboratory specializes in the isolation and characterization of these fastidious critters. During the last decade, we have isolated, characterized, and named twelve novel anaerobic fungal genera (out of the twenty-two genera currently known). We have also developed clear criteria required for the characterization and rank assignment of these fungi and have been undertaking a comprehensive multi-phasic approach in establishing a clear, Linnean-based taxonomic framework for the phylum Neocallimastigomycota. Nevertheless, we still recognize the vast diversity of AGF that are yet to be brought to culture. We continue to isolate novel anaerobic fungal taxa from a wide range of herbivores, and continue to devise novel strategies, approaches, and methodological improvements for the isolation and storage of novel AGF taxa. To view a publication from our lab on this topic, click here.
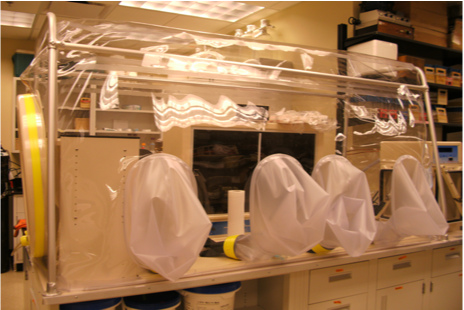
This image shows the anaerobic chamber utilized by our lab to begin the isolation process of AGF.
Patterns, determinants, and extent of the global anaerobic fungal mycobiome. In addition to culturing, we use multiple culture-independent strategies to assess AGF diversity in nature. We developed and continue to develop multiple phylogenetic markers for utilization in AGF diversity surveys. Our recent survey of AGF diversity involved a wide range of hosts from different countries and lifestyles to document patterns of occurrence, numbers, diversity, and community structure of AGF on a global scale. Our extensive surveys allowed us to test hypotheses regarding host-fungal evolutionary relationships (phylosymbiosis), as well as the role of diet, biogeography, and domestication status in shaping AGF community structure. To view a publication from our lab on this topic, click here.
Novel hosts for the Neocallimastigomycota. AGF are routinely discovered and characterized from placental mammalian herbivores. However, herbivory is known to exist long before the evolution of placental mammals. We have recently obtained tantalizing evidence on the occurrence of AGF in novel non-mammalian herbivores, e.g. marsupials and reptiles. Beyond documenting their occurrence, we are interested in studying the unique characteristics of these novel types of AGF, as well as their special adaptations and unique evolutionary trajectories in non-mammalian hosts. We are also attempting to understand the evolutionary drivers, significance, and implications of AGF and non-mammal associations.
Neocallimastigomycota evolution. We are interested in elucidating how, why, and when animals acquired and established a nutritional symbiosis with the AGF. Based on currently available data, AGF evolution as a distinct phylum appeared to have occurred relatively recently (~67 Mya) and coincided with the rise of mammalian herbivory in the late Paleogene. This estimate is far removed from the time of divergence of AGF from their closest known fungal ancestor (the chytrids), an event that occurred in the late Ediacaran (~560 Mya). This pattern implies the existence of yet-undiscovered extinct or extant taxa. By combining culture-independent diversity surveys, culturing work, genomic sequencing, phylogenomic analysis, and molecular clock timing using fossil calibration, we are trying to fill this gap in Neocallimastigomycota evolution and identify these yet-unknown lineages. We are currently testing multiple hypotheses regarding the order, timing, and interplay between two main ecological transitions (transition from aerobic to anaerobic lifestyles and transition from free-living to host-associated lifestyle) linked to Neocallimastigomycota diversification from chytrids. We also correlate these events with the body of available knowledge on animal host and substrate (plant) evolution, as well as prevalent conditions on the earth's surface and atmosphere within this timeframe.
An -omics catalog for the Neocallimastigomycota. We currently possess the world’s largest collection of AGF in pure culture covering most known AGF genera and families. We are undertaking an ambitious genomic, transcriptomic, and proteomic effort to describe the genetic landscape of the Neocallimastigomycota. We are using such information to examine dynamics of organismal and trait evolution, gene gain/loss dynamics, comparative cellulosomal production capacities, capacity for plant biomass degradation, epigenetic regulation, secondary metabolites and antibiotics production capacities, viral occurrence, oxygen detoxification machinery, potential for sexual reproduction, and the production of titan cells/survival structures in the Neocallimastigomycota.
Environmental Genomics
Zodletone Spring is a sulfide and sulfur-rich spring in southwestern Oklahoma. We have been actively researching the diversity of Zoldetone Spring for almost two decades. Due to its unique geochemistry and environmental gradients, spatial sampling in Zodletone Spring can substitute for inaccessible temporal time scales. Our current research is focusing on using the spring as a portal to characterize organisms that thrived on Earth prior to the great oxygenation event. To view a publication from our lab on this topic, click here.
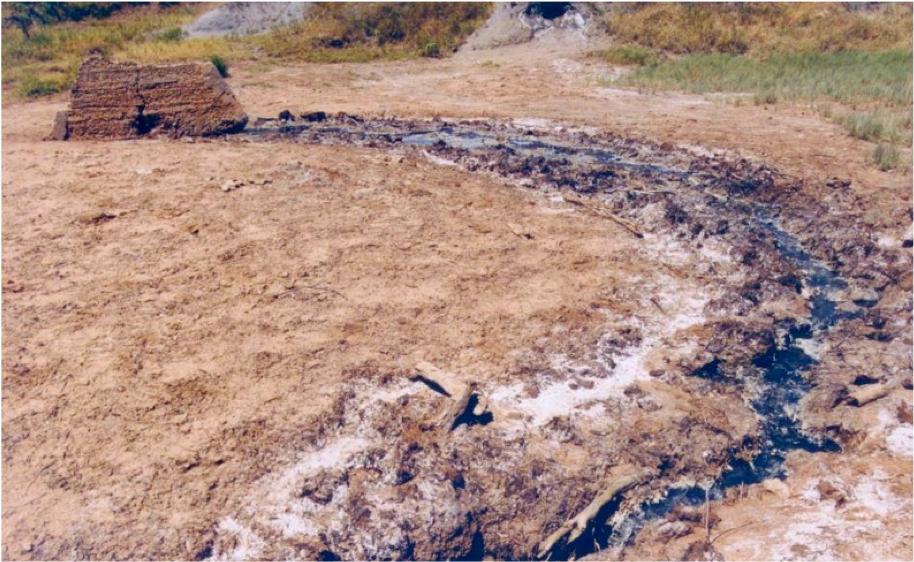

Zodletone Spring
Specifically, we are using meta-omics approaches to identify and characterize novel bacterial lineages prevalent in Zodletone Spring. In samples representing “ancient” pre-oxygenated geological eons, we have uncovered an unprecedented level of diversity and a unique microbial community dominated by taxa that are extremely rare in various current biomes on Earth. Further, we have discovered that many of these novel genera are involved in a unique sulfur cycle in the spring using precursors that are rare nowadays, but prevalent in the spring, as well as on ancient, pre-oxygenated earth (e.g. sulfite, thiosulfate, tetrathionate, and sulfur). We continue to characterize this unique community in Zodletone Spring using state-of-the-art genomic, transcriptomic, proteomic, and culturomic approaches.
Currently, our focus is on:
- Methodological improvements of genome recovery from metagenomes.
- Assessing the viriome landscape in Zodletone Spring
- Assessing novel secondary metabolites production capacity in unique members of the microbial community in Zodletone Spring.
- Utilization of novel culture-based (culturomics) approaches to isolate and characterize novel microbial taxa in the spring
A phylocentric strategy for characterizing yet-uncultured microbial taxa. Culture-independent diversity surveys have clearly shown that a large swath of microbial lineages on Earth remains uncultured. During the last decade, our laboratory has been active in using various environmental genomics and bioinformatic approaches in elucidating the metabolic role of various lineages. We use a phylocentric strategy where genomes belonging to the same lineage are characterized to provide a pangenomic view of the lineage’s collective characteristics and capacities. Such a strategy also allows for examining the relative role played by niche versus phylogeny in shaping microbial genomes and metabolic abilities. Our emphasis is on lineages thriving in anaerobic settings. Prior efforts have examined members of the phyla OP11 (Patesibacteria), WS3 (Latescibacteria), OP8 (Aminicenantes), OP1 (Bipolaricaulota), LCP89, UBP10 (Binatota), Myxococcota, Desulfobacterota, Krumholzibacterota, Mcinerneybacteria, and the archaeal phylum Diapherotrites. Current projects are focusing on characterizing the genomic content of novel lineages within the candidate phyla radiation (CPR) group and the phylum Acidobacteriota.
Freshwater Algal Blooms
Harmful algal blooms in freshwater lakes. Human activities and climate change are affecting freshwater lakes. Eutrophication and warming often lead to algal growth and changes in the lake phytoplankton and microbial community composition. One such change is the occurrence of Cyanobacterial blooms. Some species of cyanobacteria can produce harmful toxins (hence the term harmful algal blooms (HABs)) that affect both the water quality, as well as the resident pelagic and benthic communities.
Long-term data is needed to elucidate patterns of occurrence and intensity of algal blooms and to develop a mechanistic understanding of the microbial community changes leading to the blooms. Lake sedimentary DNA can be helpful in revealing historical trends of microbial community development.
We study algal bloom formation dynamics in Grand Lake, a large (18,800 ha) freshwater lake in northeastern Oklahoma. To this end, we are using current and historic sediment cores for the extraction of sedimentary DNA (sedDNA) and sedimentary ancient DNA (sedaDNA). We are undertaking an extensive microbial community characterization analysis using multiple-omics approaches, and coupling this information with historic meteorological and limnological records to understand the how, when, and why of algal bloom formation. We hope to integrate these diverse data streams using artificial intelligence and machine learning approaches to develop a predictive framework for algal bloom formation.
